iVirus is supported by
indispensable resources providing
free access for researches
iVirus integrates apps and tools, provides datasets, and designs protocols leveraging the CyVerse Cyberinfrastructure and KBase's software and data science platform. The apps iVirus develops exist within CyVerse or KBase, the datasets iVirus users use is on CyVerse or KBase's severs, and the iVirus user-guides (i.e. protocols) all use either CyVerse or KBase. These two resources are free for researchers and allow the community to analyze datasets that they might not otherwise had the capability of doing so.

CyVerse is a National Science Foundation-funded effort whose mission is "to design, deploy, and expand a national Cyberinfrastructure for Life Sciences research, and to train scientists in its use." CyVerse democratizes access to supercomputing capabilities and powerful computational infrastructure to handle huge datasets and complex analyses, enabling data-driven discovery. CyVerse is led by the University of Arizona and partners with the Texas Advanced Computing Center and Cold Spring Harbor Laboratory

In the Discovery Environment ("DE") is the interface many users will use when interacting with CyVerse. From the DE, users can find apps, view analyses, explore data, and navigate the various other resources available in CyVerse.
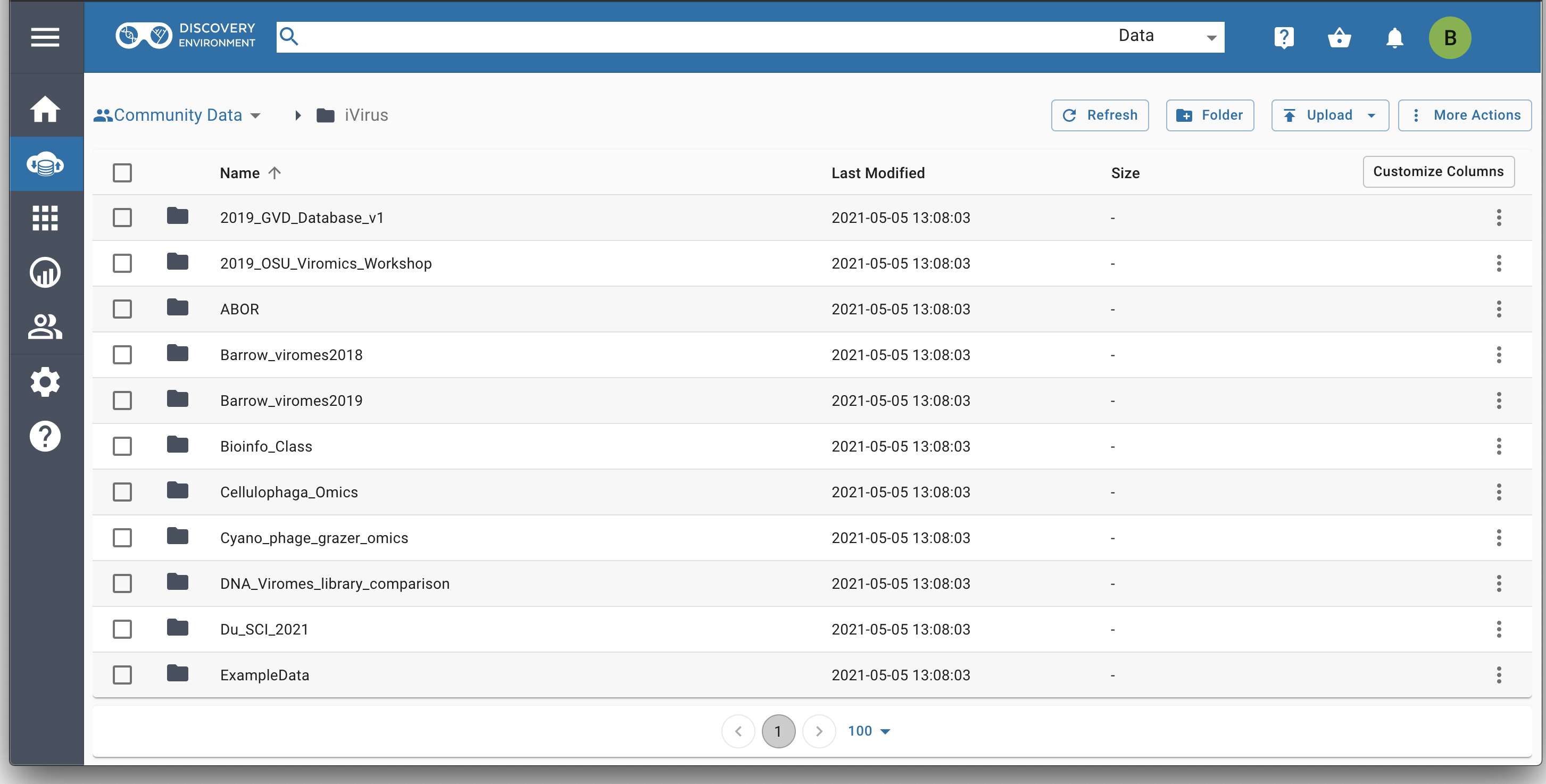
The data store is where data from analyses can be found, where other users can share data, or where community projects (such as iVirus) can publicly share their data.
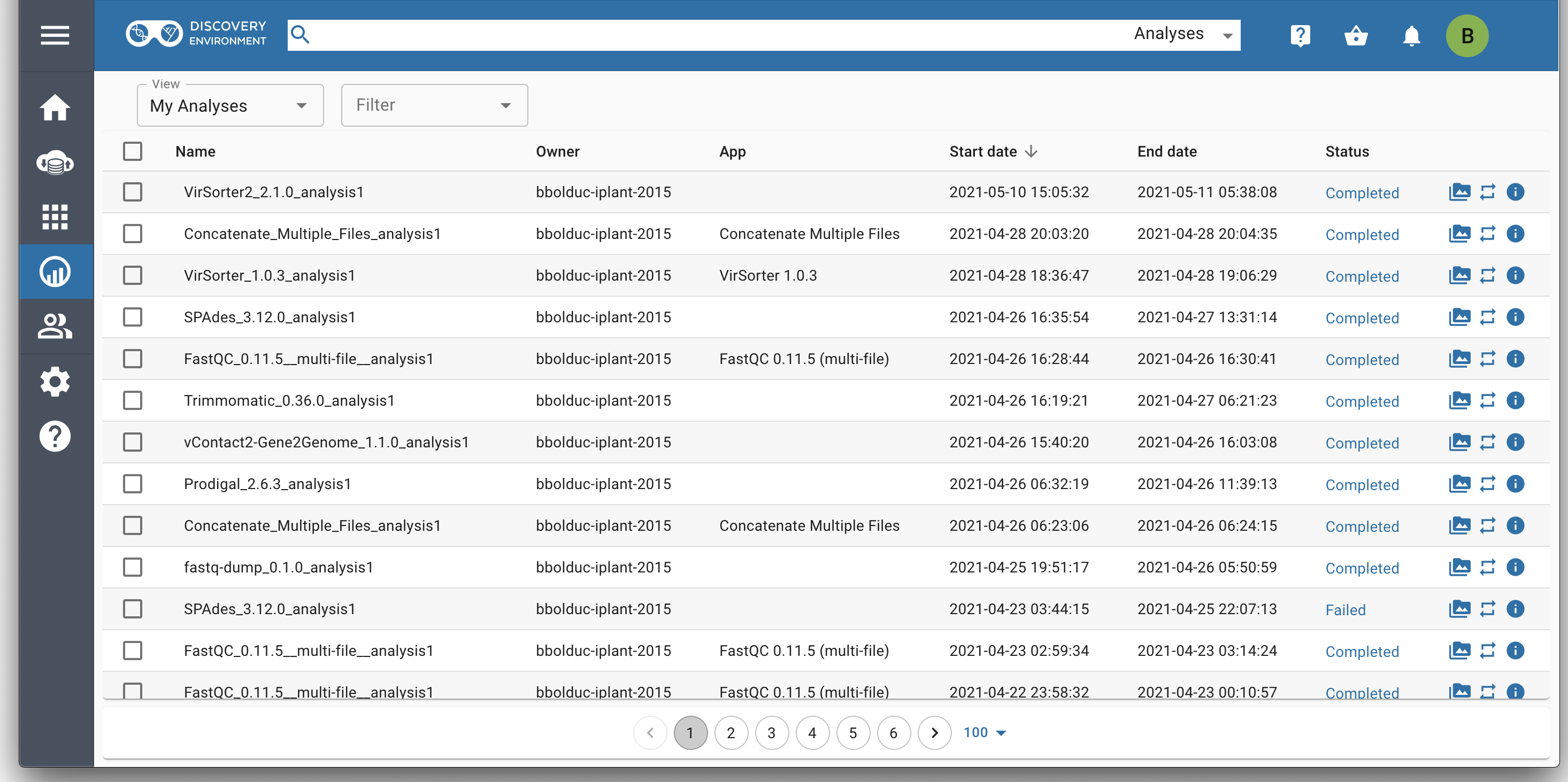
CyVerse hosts thousands of apps. CyVerse staff, along with many 3rd-party developers (e.g. external developers such as iVirus) contribute by integrating invaluable, community-focused tools.
Citing CyVerse: Anyone who uses iVirus tools and data on CyVerse is strongly encouraged to cite CyVerse:
This material is based upon work supported by the National Science Foundation under Award Numbers DBI-0735191, DBI-1265383, and DBI-1743442. URL: www.cyverse.org
For additional information, please check CyVerse's citation page.

The Department of Energy Systems Biology Knowledgebase (KBase) is a DOE-funded software and data science platform seeking to "meet the grand challenge of systems biology: predicting and designing biological function." Users can analyze, share, and collaborate using data and tools within KBase. KBase is led by Lawrence Berkeley National Laboratory with participation from Argonne, Brookhaven, and Oak Ridge National Laboratories.

KBase offers a unique take on the lab notebook, known as Narratives. These are bioinformatic workflows with provenance (i.e. data object manipulations explaining what has happened to that object) that are sharable (or can be kept private!). These Narratives can be further annotated by the user (or collaborators) to keep track of non-app-based parameters or provide background within the notebook.

Browse nearly 250 apps, manually curated by KBase staff. Apps range from read processing, genome assembly and annotation, sequence analysis, comparative genomics and microbial communities. KBase also includes metabolic modeling and expression tools, like flux-balance analysis and mass balances.

KBase can import data automatically from numerous NCBI databases and from JGI. Additionally, KBase provides apps allowing users to download directly from SRA, FTP links, and web links.
Citing KBase: Anyone who uses iVirus tools and data on KBase is strongly encouraged to cite KBase:
Arkin AP, Cottingham RW, Henry CS, Harris NL, Stevens RL, Maslov S, et al. KBase: The United States Department of Energy Systems Biology Knowledgebase. Nature Biotechnology. 2018;36: 566. doi: 10.1038/nbt.4163
For additional information, please check KBase's acknowledgement page.